The Teen Detectives Using DNA to Stalk Invasive Fish
Aquatic forensics might help target eradication efforts before things get out of control.
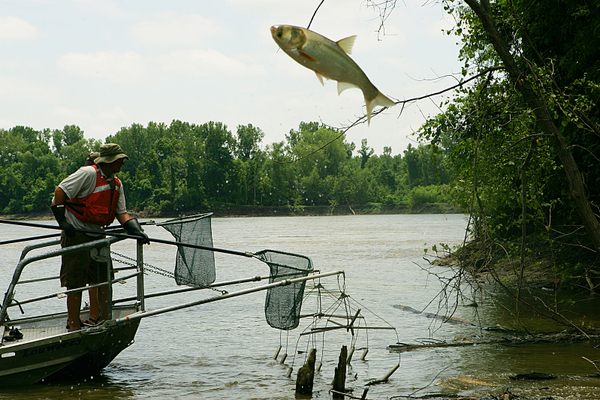
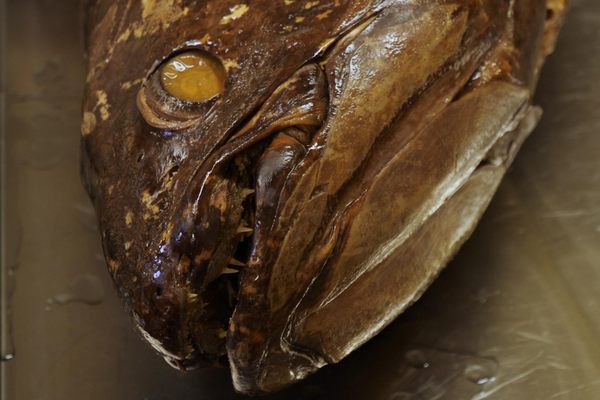
Once they infiltrate bodies of water, northern snakeheads are hard to shake. The invasive fish lock their needle-sharp teeth around other fish, native birds, and small mammals, and they reproduce like crazy. Within two years, a single female can deposit 150,000 eggs. As long as they stay wet, they can survive on land for days, flailing their fins and tails to propel themselves along, from one body of water to the next.
So far, New York State has two outbreaks of the unwelcome guests, which are native to Asia—one that has been wrestled under control, and one with a much more problematic prognosis. Left unchecked, the population in a lake in Orange County “has the potential to infest the entire Hudson River drainage and beyond to the Great Lakes and continental U.S.,” according to the Department of Environmental Conservation.
By the time an invasive species fully colonizes a new habitat, it’s nearly impossible to completely eradicate. “To be perfectly honest, that almost never works,” says Donna Cassidy-Hanley, a senior research associate at Cornell’s College of Veterinary Medicine. An alien population can be managed before it reaches a critical mass—but that’s also the time when it’s most difficult to detect and track. New York State is shot through with thousands of ponds and lakes. It’s not feasible to wade into all of them, eyes peeled for a flash of the snakehead’s saddle-patterned scales. So, on a drizzly fall morning, a huddle of students set out to look a different way.

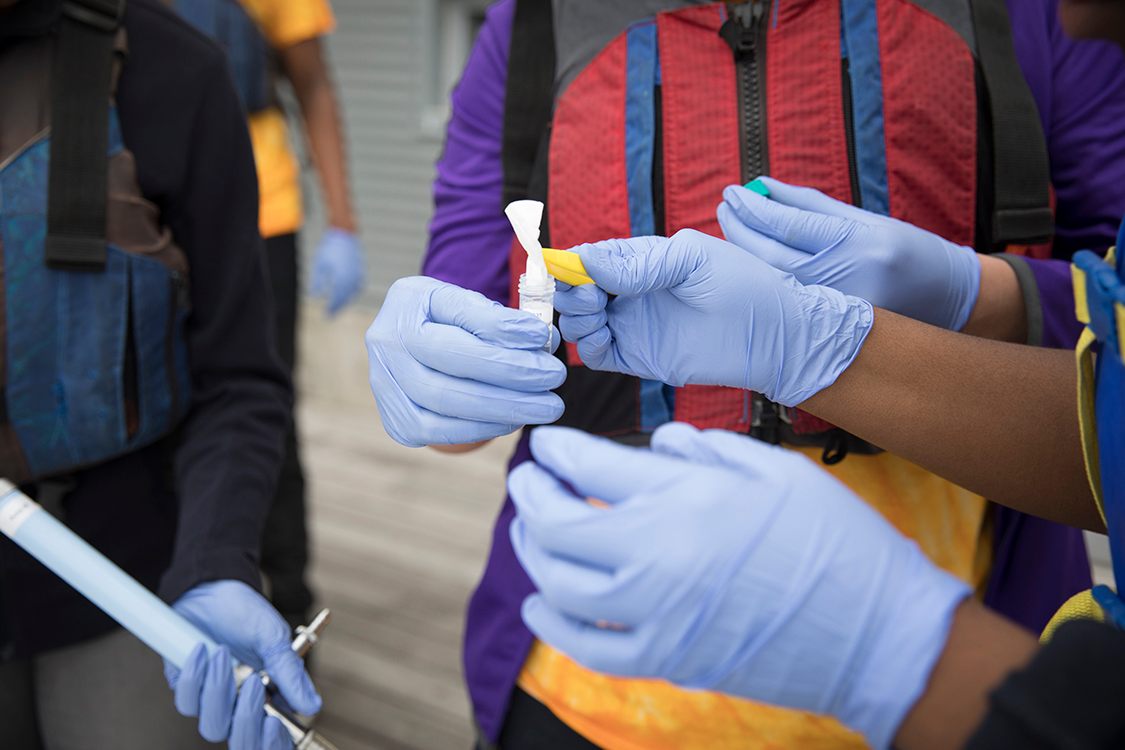
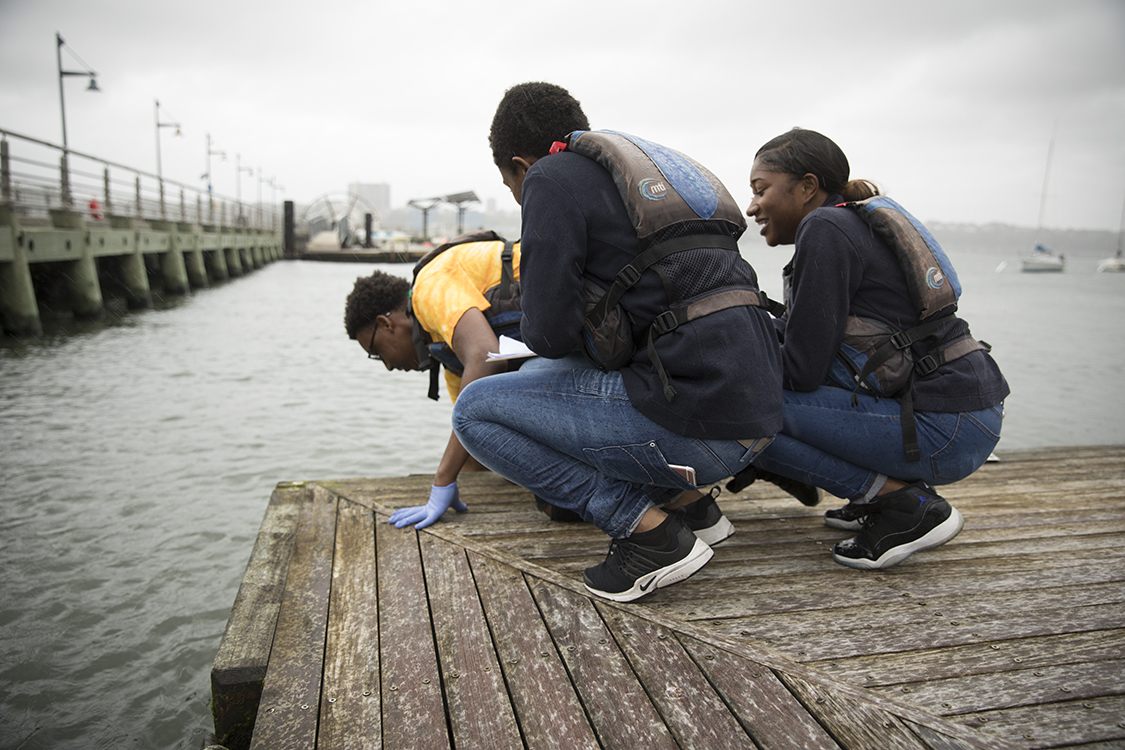
Through Cornell’s FishTracker program, funded by the USDA National Institute of Food and Agriculture, teams of life-jacket-clad teenagers leaned out over docks to scoop water samples from the Hudson River. They plunged plastic bags into the water column, trying to avoid both the surface muck and mud stirred up from the bottom. They then stabbed each bag with a toothpick, and let the trickle drip through a filter, which went back to the lab.
What each filter captures is a jumble of genetic material floating around in the water, sloughed off of the creatures that call the river home, or even just passed through. In the lab, researchers use a technique called quantitative PCR (qPCR), which lets them target specific DNA signatures in the sample. “The technology is right off the shelf,” says James Casey, a virologist at the College of Veterinary Medicine, who developed the genetic tests used in the study. “Every fish in the Great Lakes and NYC has been barcoded, just like the universal product code that you scan in the supermarket.” Using these distinctive genetic identifiers, Casey says, “we can make primers and probes that will specifically identify only that.” For now, the FishTracker project is looking out for a range of alien occupiers of American waters: sea lampreys, Asian carp, round goby, northern snakehead, white perch, and Asian swamp eel. Under ideal conditions, the technology can suss out a single fish in a 50-acre pond that is 10 feet deep. And because DNA hangs in the water for up to six hours, the test can even flag visitors that have moved on.
Since the pilot program kicked off three years ago, more than 3,000 students from 78 schools have surveyed 320 sites across New York State. They’re currently monitoring for six invasive species and two endangered ones, the American eel and the deepwater cisco. For students, the program is an accessible, up-close introduction to otherwise opaque ideas about ecology, genomics, and conservation. “It brings them into science in a real sense,” Casey says. “It starts a conversation that students can’t get away from.” And since students pair their samples with GPS coordinates, the data—which are mapped and available on an open-access website—can tell help researchers and partners decide where to focus tailored eradication efforts.

Each of these DNA-laden filters is a snapshot of a particular ecosystem at a certain moment, so the data they contain are still valuable, even after they have been scanned for invasive DNA. The researchers are creating a stockpile of them, kept at -80 degrees Celsius, so they can be screened again later on for some other species, or to see how changes such as climate and development alter ecosystems, driving some species away from their current habitats or giving others room to flourish.
As far as actually stopping these invasives from taking over, the work is “a drop in the bucket,” Cassidy-Hanley says, but it’s “establishing a mechanism that is expandable, dependable, and relatively economic.” And it gives kids a taste of aquatic forensic science in the process.













Follow us on Twitter to get the latest on the world's hidden wonders.
Like us on Facebook to get the latest on the world's hidden wonders.
Follow us on Twitter Like us on Facebook